Difference between revisions of "Main Page"
From Zhang Laboratory
| (185 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
{{#meta: keywords | computational biology; systems biology; RNA splicing and regulatory networks; gene expression }}{{#meta: description | The Chaolin Zhang Laboratory Home Page at Columbia University}}{{#meta: Content-Type | text/html; charset=utf-8 }} | {{#meta: keywords | computational biology; systems biology; RNA splicing and regulatory networks; gene expression }}{{#meta: description | The Chaolin Zhang Laboratory Home Page at Columbia University}}{{#meta: Content-Type | text/html; charset=utf-8 }} | ||
| − | [[File: | + | <div style='text-align: right;'> |
| + | [[File:Twitter.jpg|25px|link=https://twitter.com/chaolinzhang]] [[File:LinkedIn.png|20px|link=https://www.linkedin.com/in/chaolin-zhang-52835217/]] | ||
| + | </div> | ||
<html> | <html> | ||
| Line 63: | Line 65: | ||
<div> | <div> | ||
| − | <a u=image href="/index.php/Research" rel="nofollow"><img u="image" src="/data/images/slideshow/deconstruct_splice_code.png" width="600" height="300" /></a> | + | <a u=image href="/index.php/Research" title="Our long term goal: better mechanistic understanding of RNA splicing regulation and function." rel="nofollow"><img u="image" src="/data/images/slideshow/deconstruct_splice_code.png" width="600" height="300" /></a> |
</div> | </div> | ||
| + | |||
| + | <div> | ||
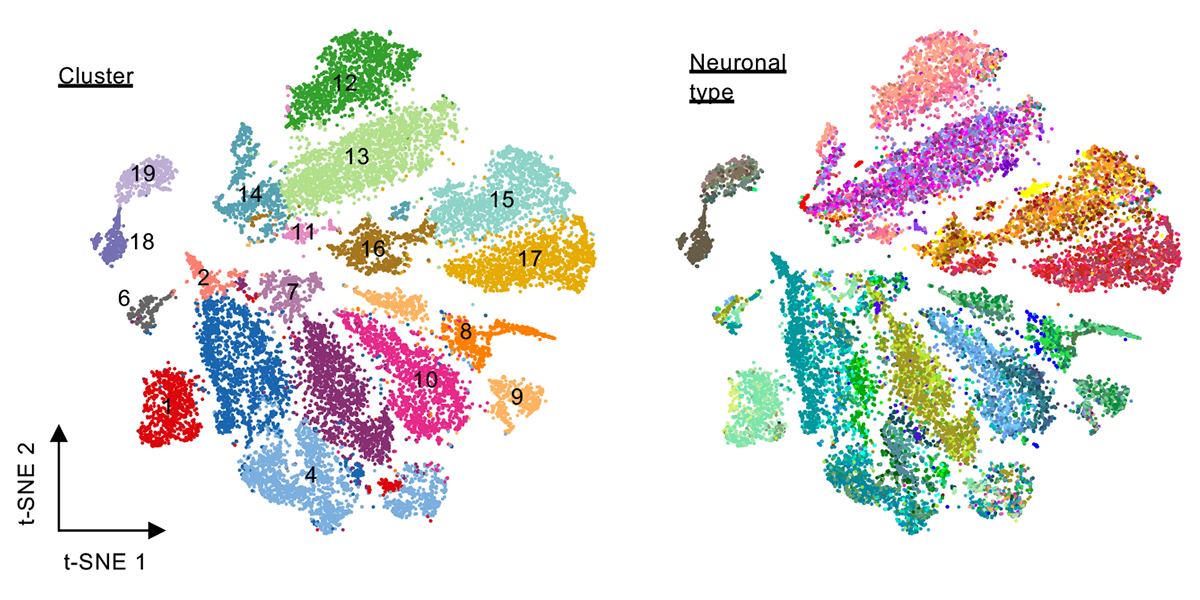
| + | <a u=image href="https://www.pnas.org/content/118/10/e2013056118/tab-article-info" title="Clustering of neuronal subtypes based on splicing profiles (PNAS, 2021)." rel="nofollow"><img u="image" src="/data/images/slideshow/neuronal_subtype_splicing.png" width="600" height="300" /></a> | ||
| + | </div> | ||
| + | |||
| + | <div> | ||
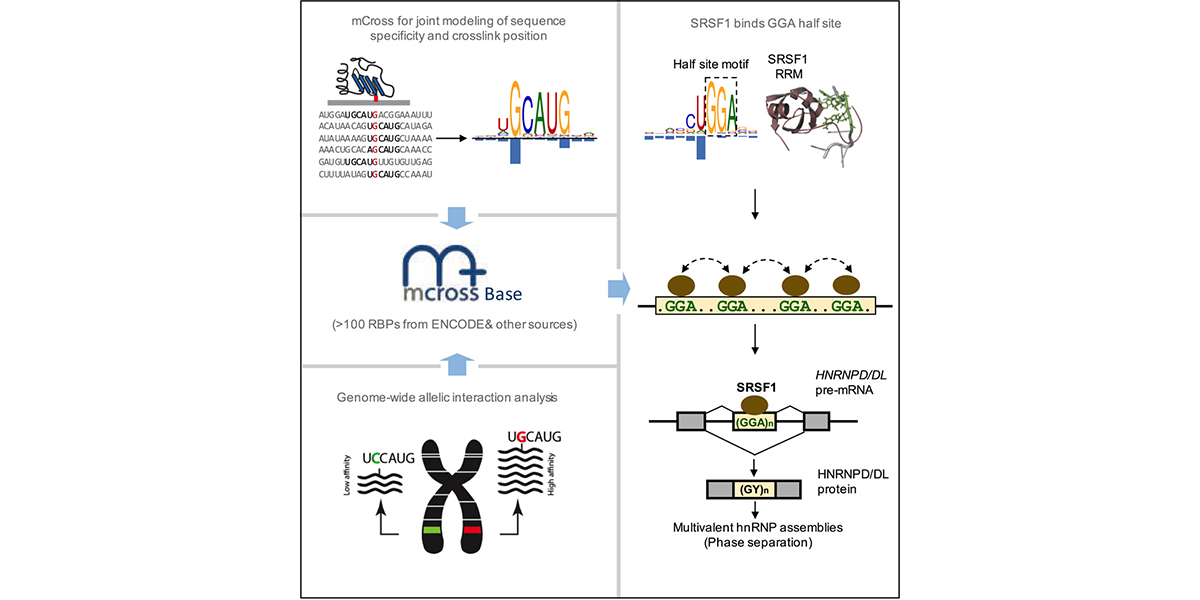
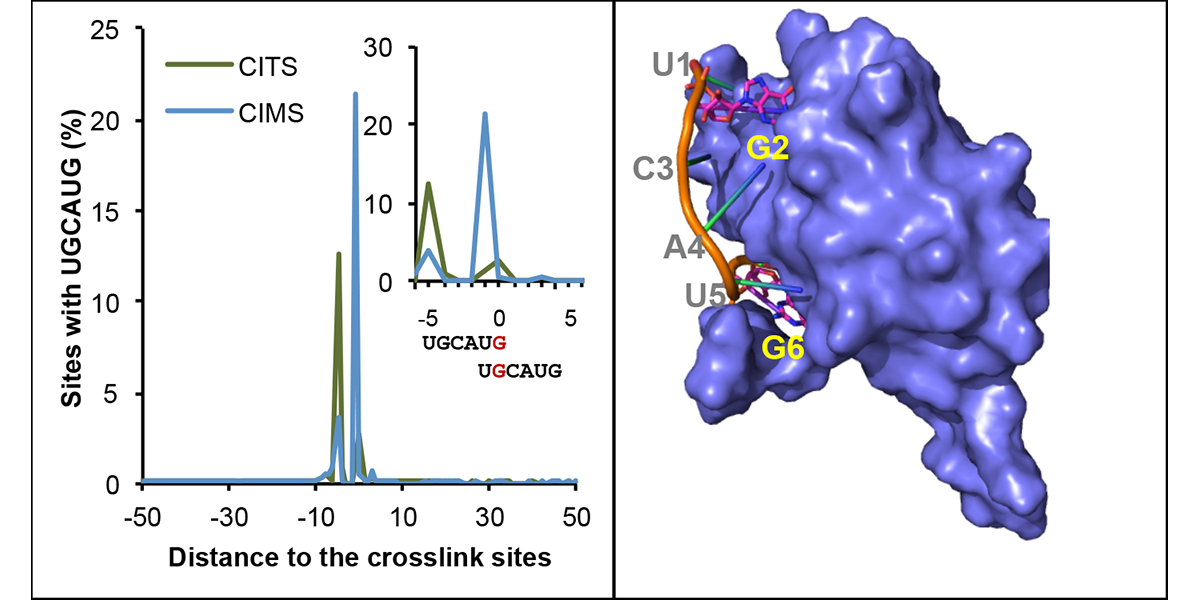
| + | <a u=image href="https://www.sciencedirect.com/science/article/pii/S1097276519300929" title="mCross: precise registration of protein-RNA crosslink sites to define RBP binding specificity using CLIP data (Mol. Cell, 2019)." rel="nofollow"><img u="image" src="/data/images/slideshow/mcross.png" width="600" height="300" /></a> | ||
| + | </div> | ||
| + | |||
<div> | <div> | ||
| − | <a u=image href="https://authors.elsevier.com/a/1XPn83vVUP60qL" rel="nofollow"><img u="image" src="/data/images/slideshow/lin28.png" width="600" height="300" /></a> | + | <a u=image href="https://authors.elsevier.com/a/1XPn83vVUP60qL" title="A subclass of let-7 microRNA partially escape from LIN28 repression (Mol. Cell, 2018)." rel="nofollow"><img u="image" src="/data/images/slideshow/lin28.png" width="600" height="300" /></a> |
</div> | </div> | ||
<div> | <div> | ||
| − | <a u=image href="https://www.nature.com/articles/s41467-018-04559-0" rel="nofollow"><img u="image" src="/data/images/slideshow/devas.png" width="600" height="300" /></a> | + | <a u=image href="https://www.nature.com/articles/s41467-018-04559-0" title="Precise timing of alternative splicing switches during neuronal development (Nat. Commun., 2018)" rel="nofollow"><img u="image" src="/data/images/slideshow/devas.png" width="600" height="300" /></a> |
</div> | </div> | ||
<div> | <div> | ||
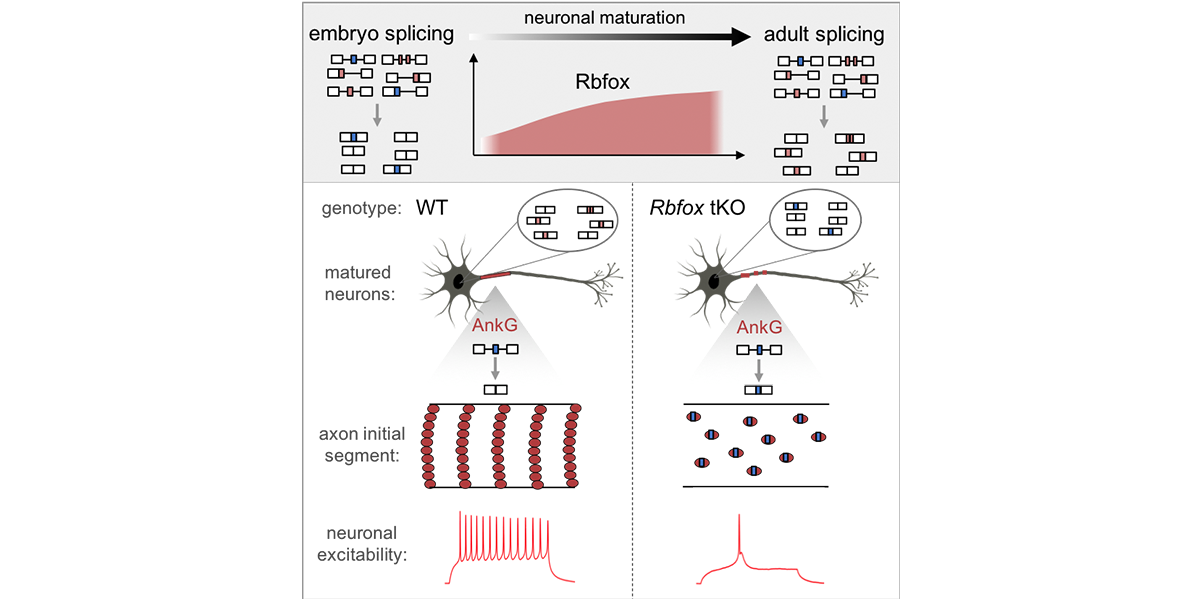
| − | <a u=image href="http://www.cell.com/neuron/fulltext/S0896-6273(18)30023-0" rel="nofollow"><img u="image" src="/data/images/slideshow/ais.jpg" width="600" height="300" /></a> | + | <a u=image href="http://www.cell.com/neuron/fulltext/S0896-6273(18)30023-0" title="Rbfox regulates axon initial segment and neuronal excitability (Neuron, 2018)." rel="nofollow"><img u="image" src="/data/images/slideshow/ais.jpg" width="600" height="300" /></a> |
</div> | </div> | ||
<div> | <div> | ||
| − | <a u=image href="http://www.cell.com/neuron/fulltext/S0896-6273(18)30023-0" rel="nofollow"><img u="image" src="/data/images/slideshow/RbfoxMN.png" width="600" height="300" /></a> | + | <a u=image href="http://www.cell.com/neuron/fulltext/S0896-6273(18)30023-0" title="Rbfox regulates axon initial segment and neuronal excitability (Neuron, 2018)." rel="nofollow"><img u="image" src="/data/images/slideshow/RbfoxMN.png" width="600" height="300" /></a> |
</div> | </div> | ||
<div> | <div> | ||
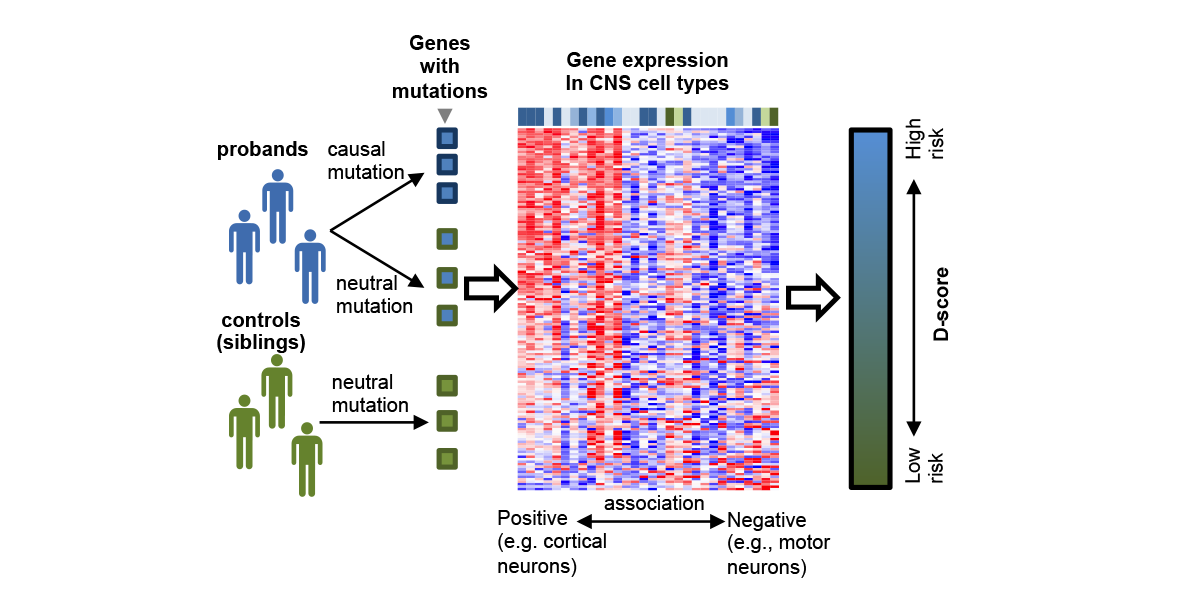
| − | <a u=image href="http://onlinelibrary.wiley.com/doi/10.1002/humu.23147/abstract" rel="nofollow"><img u="image" src="/data/images/slideshow/DAMAGES.png" width="600" height="300" /></a> | + | <a u=image href="http://onlinelibrary.wiley.com/doi/10.1002/humu.23147/abstract" title="A neuronal-cell type-specific gene expression signature predicts autism risk genes (Hum. Mut. 2017)." rel="nofollow"><img u="image" src="/data/images/slideshow/DAMAGES.png" width="600" height="300" /></a> |
</div> | </div> | ||
<div> | <div> | ||
| − | <a u=image href="/index.php/Research" rel="nofollow"><img u="image" src="/data/images/slideshow/dual_splice_site.png" width="600" height="300" /></a> | + | <a u=image href="/index.php/Research" title="Genes have 5' splice sites and 3' splice sites, but some splice sites have dual roles" rel="nofollow"><img u="image" src="/data/images/slideshow/dual_splice_site.png" width="600" height="300" /></a> |
</div> | </div> | ||
<div> | <div> | ||
| − | <a u=image href="/index.php/Research" rel="nofollow"><img u="image" src="/data/images/slideshow/crosslink.png" width="600" height="300" /></a> | + | <a u=image href="/index.php/Research" title="UV-crosslinking induced protein-RNA adducts provide a signature to determine protein-RNA interactions at single nucleotide resolution" rel="nofollow"><img u="image" src="/data/images/slideshow/crosslink.png" width="600" height="300" /></a> |
</div> | </div> | ||
<div> | <div> | ||
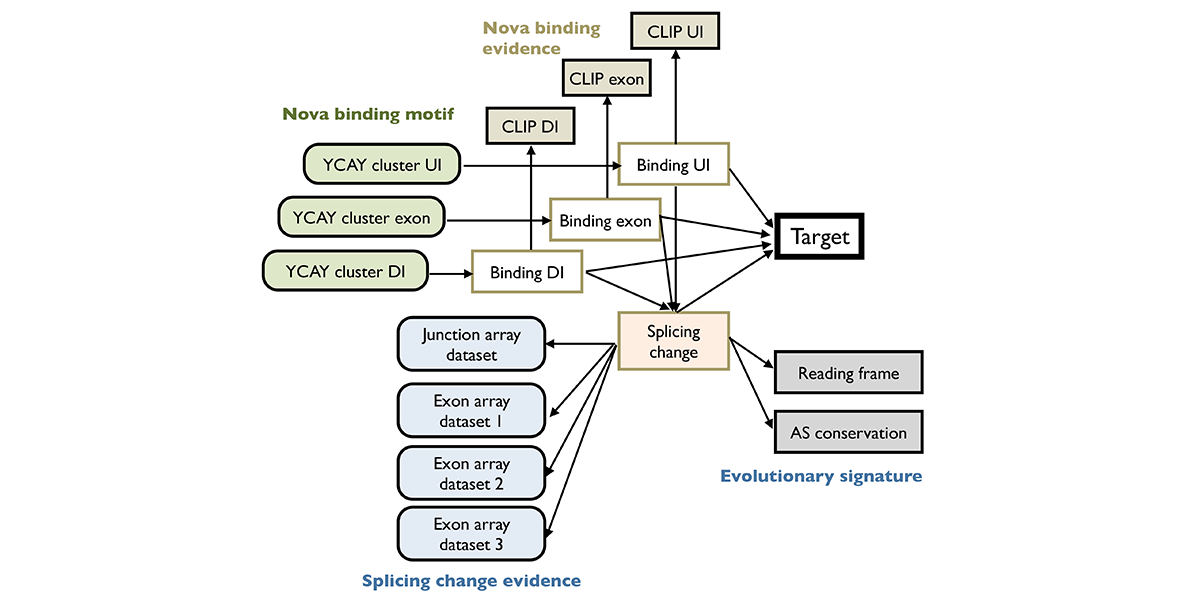
| − | <a u=image href="/index.php/Research" rel="nofollow"><img u="image" src="/data/images/slideshow/bnet.png" width="600" height="300" /></a> | + | <a u=image href="/index.php/Research" title="An integrative modeling approach to construct splicing regulatory networks" rel="nofollow"><img u="image" src="/data/images/slideshow/bnet.png" width="600" height="300" /></a> |
</div> | </div> | ||
<div> | <div> | ||
| − | <a u=image href="/index.php/Research" rel="nofollow"><img u="image" src="/data/images/slideshow/brain_dev.png" width="600" height="300" /></a> | + | <a u=image href="/index.php/Research" title="We aim to understand the contribution to alternative splicing to transcriptome diversity in brain development" rel="nofollow"><img u="image" src="/data/images/slideshow/brain_dev.png" width="600" height="300" /></a> |
</div> | </div> | ||
<div> | <div> | ||
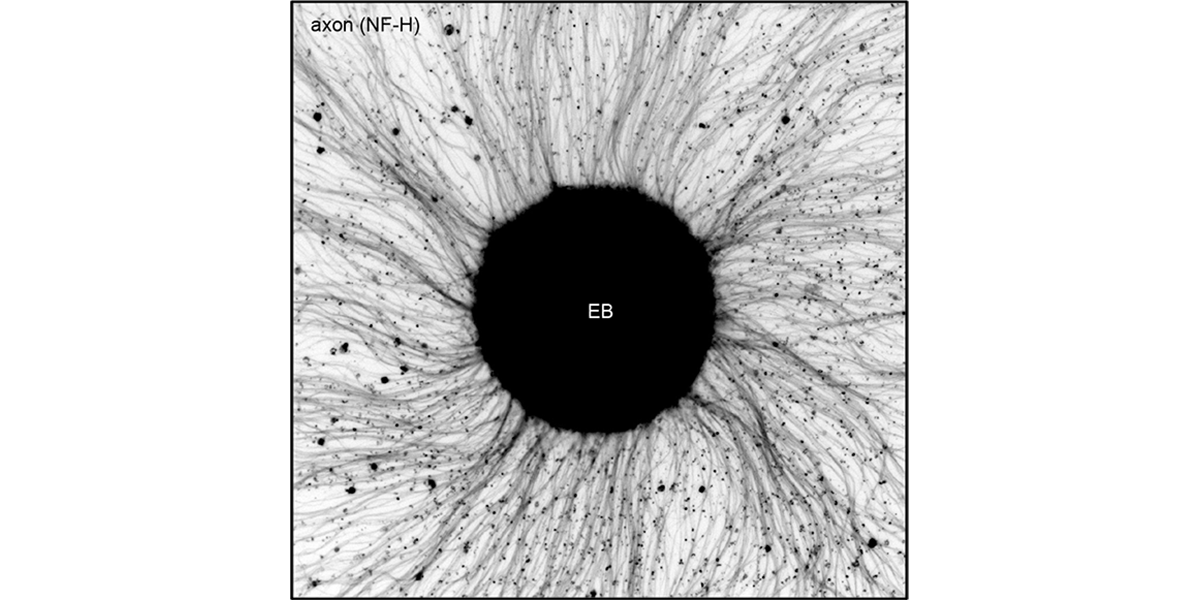
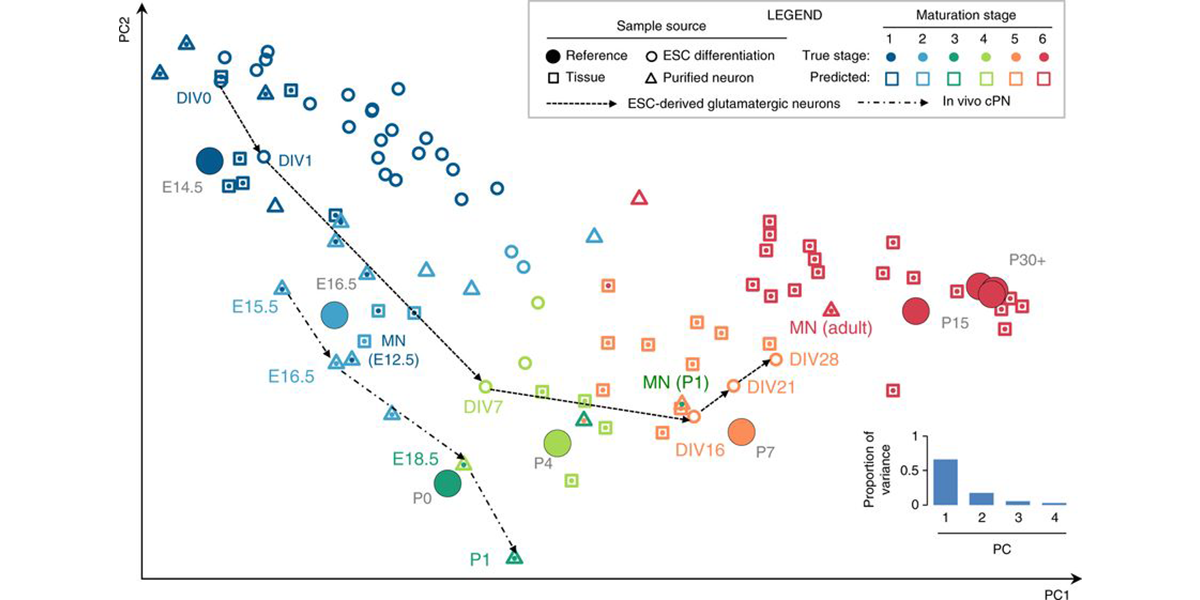
| − | <a u=image href="/index.php/Research" rel="nofollow"><img u="image" src="/data/images/slideshow/eb.png" width="600" height="300" /></a> | + | <a u=image href="/index.php/Research" title="ESC-derived motor neuronsprovide an ideal system to model neural development and maturation in vitro" rel="nofollow"><img u="image" src="/data/images/slideshow/eb.png" width="600" height="300" /></a> |
</div> | </div> | ||
| Line 122: | Line 133: | ||
| − | + | ||
| + | ==Introduction of the Zhang Laboratory== | ||
| + | |||
| + | {|class="zebra wikitable" width="100%" style="border:1px solid; border-collapse:collapse;" | ||
| + | |||
| + | |- | ||
| + | |style="width: 1px"| | ||
We are part of the [http://systemsbiology.columbia.edu Department of Systems Biology], the [http://cpmcnet.columbia.edu/dept/gsas/biochem/ Department of Biochemistry and Molecular Biophysics], and the [http://www.columbiamnc.org/ Motor Neuron Center] at [http://http://www.cumc.columbia.edu Columbia University Medical Center]. | We are part of the [http://systemsbiology.columbia.edu Department of Systems Biology], the [http://cpmcnet.columbia.edu/dept/gsas/biochem/ Department of Biochemistry and Molecular Biophysics], and the [http://www.columbiamnc.org/ Motor Neuron Center] at [http://http://www.cumc.columbia.edu Columbia University Medical Center]. | ||
| Line 128: | Line 145: | ||
We are fascinated by the complexity of the mammalian nervous system and its underlying molecular mechanisms. While mammals have a similar number of genes compared to phenotypically simpler organisms (such as worms), one apparent feature of mammalian genes is their more complicated gene structures, providing an opportunity for sophisticated regulation at the RNA level. | We are fascinated by the complexity of the mammalian nervous system and its underlying molecular mechanisms. While mammals have a similar number of genes compared to phenotypically simpler organisms (such as worms), one apparent feature of mammalian genes is their more complicated gene structures, providing an opportunity for sophisticated regulation at the RNA level. | ||
| − | The focus of the Zhang lab is to dissect RNA regulatory networks in the nervous system as a way to understand the mammalian complexity manifested in evolutionary-developmental (evo-devo) processes and in several neuronal disorders. | + | The focus of the Zhang lab is to dissect RNA regulatory networks in the nervous system as a way to understand the mammalian complexity manifested in evolutionary-developmental (evo-devo) processes and in several neuronal disorders. The lab has a mixed dry and wet setup, and takes a multidisciplinary approach that tightly integrates high-throughput biochemistry, genomics and computational approaches applied to mouse models and in vitro neuronal differentiation systems from pluripotent stem cells. On the mechanistic side, the Zhang lab focuses on fundamental understanding of the targeting specificity of RNA-binding proteins (RBPs), how they regulate alternative splicing in various cellular contexts, especially in the nervous system, and how such regulation can be disrupted by mutations and genetic variations. On the functional side, the lab aims to uncover the roles of RBPs in determining the neuronal cell fate, morphological and functional properties during neural differentiation and maturation. The lab has also been working on translating fundamental knowledge on RNA regulation to precision genetic medicine, with a particular focus on multiple devastating monogenic diseases affecting the central nervous system. |
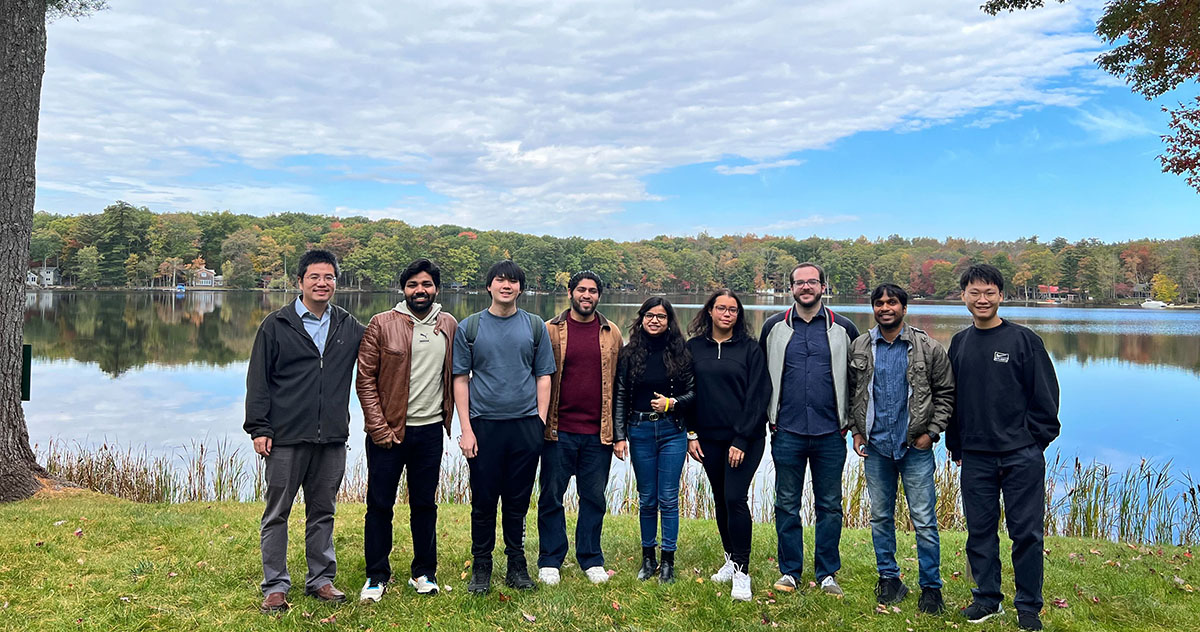
| − | + | [[File:DSBRetreat2023s2.jpg|920px|center|link=]] | |
| − | + | ||
| − | + | ||
| − | + | ||
| + | <div style="text-align:center"> | ||
| + | 2023 Systems Biology Department Retreat (Woodloch Pines Resort, PA) | ||
| + | </div> | ||
| + | |} | ||
| + | |||
| + | |||
| + | ==Lab News== | ||
| + | |||
| + | {| | ||
| + | |- | ||
| + | |style="width: 50%;vertical-align:top| | ||
| + | |||
| + | [[File:CSHA.jpg|600px|link=https://www.csh-asia.org/?content/2449]] | ||
| + | |||
| + | Chaolin Zhang will co-organize the 2024 Cold Spring Harbor Asia meeting "Computational Biology of the Genome", held in Suzhou China on Oct 21-25. | ||
| + | |||
| + | |style="width: 50%;vertical-align:top| | ||
| + | |||
| + | * 03/26/2024. Preprint release - [http://www.biorxiv.org/content/10.1101/2024.03.22.586363v1 DeltaSplice] for quantitative prediction of splicing and splicing-altering mutations. | ||
| + | * 03/22/2024. Congrats Yocelyn Recinos, recent graduate from the lab, for winning the [http://systemsbiology.columbia.edu/news/yocelyn-recinos-recipient-of-the-2024-dean’s-award-for-excellence-in-research 2024 Dean's Award for Excellence in Research]. | ||
| + | * 03/13/2024. Our PxR3D-map paper is officially out in Nature Communications. http://www.nature.com/articles/s41467-024-46429-y | ||
| + | * 02/20/2024. Welcome Shane Chu and Ye Wang, who joined the Zhang lab as postdoc recently. | ||
| + | * 11/22/2023. Congrats Dan Moakley for successfully defending the PhD thesis on neuron type-specific alternative splicing regulation! | ||
| + | * 10/17/2023. Congrats Yocelyn Recinos for winning the poster award at Systems Biology department retreat! | ||
| + | * 10/02/2023. (Belated news) We released our [http://www.biorxiv.org/content/10.1101/2023.08.21.554109v1 preprint] on SpliceRUSH, a high-throughput screening method for mapping both proximal and distal splicing-regulatory elements in native sequences. Congrats Yocelyn and team! | ||
| + | * 10/02/2023. (Belated news) Welcome new PhD students Albertine Albertine Neal, Fahad Paryani, and Tianji Yu for rotating in the lab! | ||
| + | * 07/17/2023. Welcome Madeline Scalon, Vivian Coraci, and Matthew Yuan for joining the lab as visiting high school students to perform their summer research! | ||
| + | * 07/14/2023. Welcome Ruchika for starting her postdoc in the lab! | ||
| + | |||
| + | |||
| + | |||
| + | <div style="text-align:right"> | ||
| + | >>> [[Lab news|all lab news]] | ||
| + | </div> | ||
| + | |} | ||
| + | |||
| + | ==Where We Are== | ||
| + | |||
| + | {|class="zebra wikitable" width="100%" style="border:1px solid; border-collapse:collapse;" | ||
| + | |||
| + | |- | ||
| + | |style="width: 1px" align="center"| | ||
| + | |||
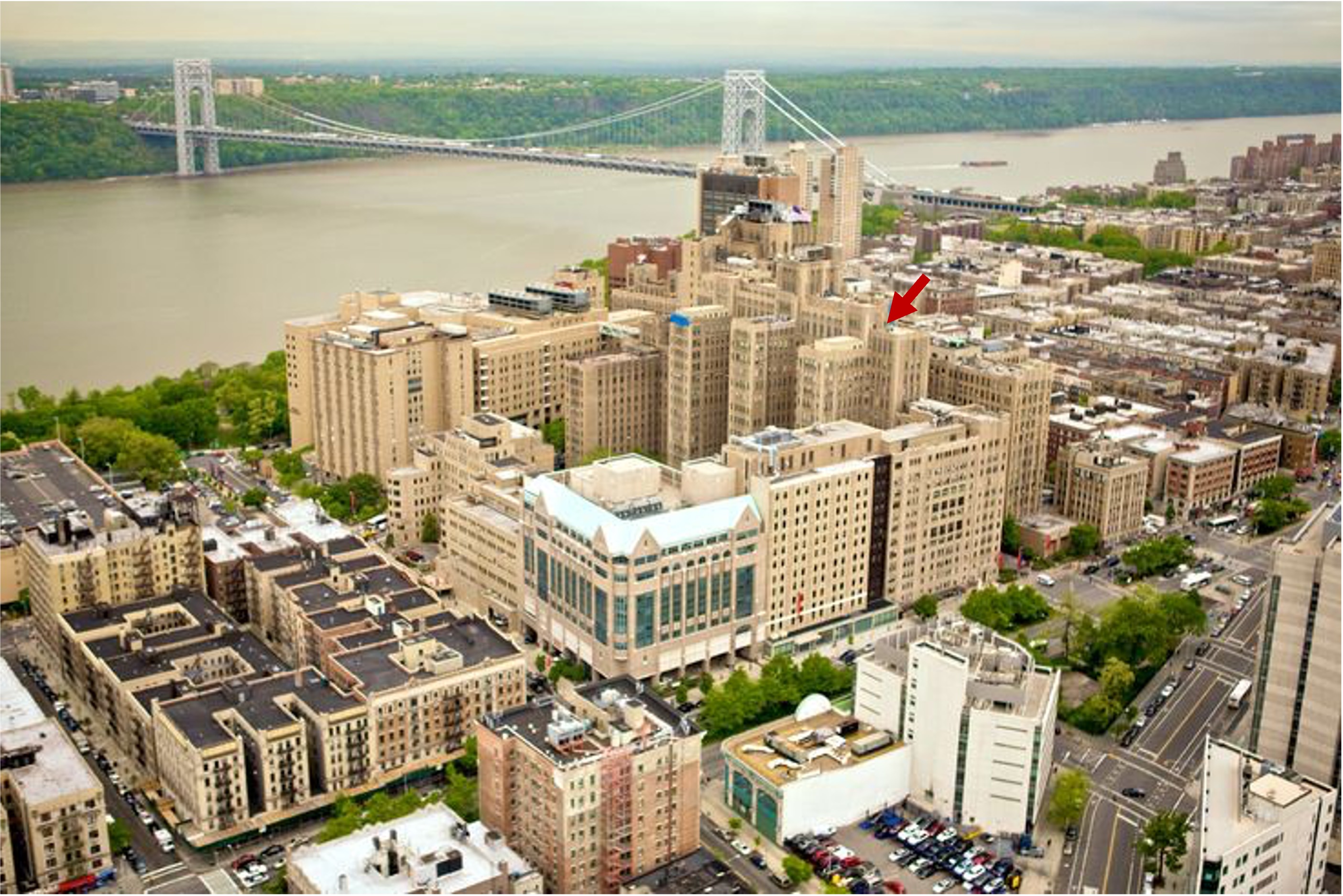
| + | [[File:Cumc2.jpg|600px|Columbia University Medical Center is located in the Washington Heights neighborhood of Manhattan in New York City. It is situated at the northern tip of Manhattan, at the intersection of West 168th Street and Broadway. The Medical Center is easily accessible by public transportation, with the 1, A and C trains stopping at the 168th Street station.|link=]] | ||
| + | |||
| + | | | ||
| + | |||
| + | [[File:Washingtonheight.jpeg|320px|link=]] | ||
| + | |||
| + | |- | ||
| + | |||
| + | | | ||
| + | <div style="text-align:center"> | ||
| + | Columbia University Irving Medical Center (image credit: [http://www.pinterest.com/pin/columbia-university-medical-center-partially-manhatten-nyc--619385754984816311/ internet]) | ||
| + | </div> | ||
| + | | | ||
| + | <div style="text-align:center"> | ||
| + | Washington Height, NYC (image credit: [http://twitter.com/KellyrKopp @KellyrKopp/twitter]) | ||
| + | </div> | ||
| + | |} | ||
<html> | <html> | ||
| + | |||
<div align="center"> | <div align="center"> | ||
| − | <a href="http:// | + | |
| + | <a href="http://systemsbiology.columbia.edu"><img src="/data/images/CSB.png" width="147" height="40" /></a> | ||
| + | <a href="http://www.biochem.cuimc.columbia.edu"><img src="/images/d/d6/Logo_biochem.jpg" width="206" height="40" /></a> | ||
<a href="http://www.columbiamnc.org/"><img src="/data/images/mnc_logo.gif" width="122" height="35" /></a> | <a href="http://www.columbiamnc.org/"><img src="/data/images/mnc_logo.gif" width="122" height="35" /></a> | ||
<a href="http://www.c2b2.columbia.edu"><img src="/data/images/C2B2_logo.png" width="215" height="60" /></a> <p> | <a href="http://www.c2b2.columbia.edu"><img src="/data/images/C2B2_logo.png" width="215" height="60" /></a> <p> | ||
Latest revision as of 10:27, 26 March 2024
Introduction of the Zhang Laboratory
|
We are part of the Department of Systems Biology, the Department of Biochemistry and Molecular Biophysics, and the Motor Neuron Center at Columbia University Medical Center. We are fascinated by the complexity of the mammalian nervous system and its underlying molecular mechanisms. While mammals have a similar number of genes compared to phenotypically simpler organisms (such as worms), one apparent feature of mammalian genes is their more complicated gene structures, providing an opportunity for sophisticated regulation at the RNA level. The focus of the Zhang lab is to dissect RNA regulatory networks in the nervous system as a way to understand the mammalian complexity manifested in evolutionary-developmental (evo-devo) processes and in several neuronal disorders. The lab has a mixed dry and wet setup, and takes a multidisciplinary approach that tightly integrates high-throughput biochemistry, genomics and computational approaches applied to mouse models and in vitro neuronal differentiation systems from pluripotent stem cells. On the mechanistic side, the Zhang lab focuses on fundamental understanding of the targeting specificity of RNA-binding proteins (RBPs), how they regulate alternative splicing in various cellular contexts, especially in the nervous system, and how such regulation can be disrupted by mutations and genetic variations. On the functional side, the lab aims to uncover the roles of RBPs in determining the neuronal cell fate, morphological and functional properties during neural differentiation and maturation. The lab has also been working on translating fundamental knowledge on RNA regulation to precision genetic medicine, with a particular focus on multiple devastating monogenic diseases affecting the central nervous system.
 2023 Systems Biology Department Retreat (Woodloch Pines Resort, PA) |
Lab News
|
Chaolin Zhang will co-organize the 2024 Cold Spring Harbor Asia meeting "Computational Biology of the Genome", held in Suzhou China on Oct 21-25. |
>>> all lab news |
Where We Are
|
|
|
|
Columbia University Irving Medical Center (image credit: internet) |
Washington Height, NYC (image credit: @KellyrKopp/twitter) |